In a 9-minute episode of The Miller Report: Real Clear Journalism, Maggie Miller sits down with Jesse Ausubel, Director of the Program for the Human Environment at Rockefeller University, to discuss the future of energy, environmental realism, and what America’s energy system could look like in 2030 and 2050.
News
Scenario bias started at the outset of the IPCC
The Intergovernmental Panel on Climate Change has recently retreated from its fantastical “8.5” scenarios, as reported in a recent column by Michael Shellenberger on the capture of the IPCC by those with an alarmist bias that cites Jesse Ausubel. The 8.5 scenario originated in the year 2000. But the Business-As-Usual scenario in the first IPCC report in 1990 was equally extreme. As a reviewer of the first IPCC report, Jesse critiqued the BAU scenario as “Brezhnevite” but his review was totally ignored.
Below are a few paragraphs that repeat Jesse’s 1990 critique of the IPCC BAU scenario from Jesse’s 1994 paper Technical Progress and Climatic Change published in Energy Policy 23 (4/5): 411–416 1995 (also pp. 501-512 in “Integrated Assessment of Mitigation, Impacts, and Adaptation to Climate Change,” N Nakicenovic, WD Nordhaus, R Richels, and FL Toth (eds), International Institute for Applied Systems Analysis, Laxenburg, Austria, 1994).
Excerpt: This possibility is illustrated by the final technological trajectory discussed here, that of decarbonization, or the decreasing carbon intensity of primary energy, measured in tons of carbon created per kilowatt year of electricity (or its equivalent) (Figure 8). As is evident, the global energy system has been steadily economizing on carbon. Without gloomy climate forecasts or dirty taxes.
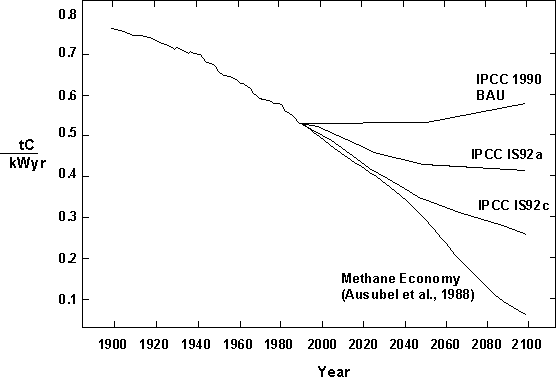
Figure 8. Carbon intensity of primary energy, 1900-1990, with projections to 2100. The projection stopping the historic trend of decarbonization is the IPCC 1990 “Business as Usual” (BAU) scenario; IPCC IS92a and ISP2c are high and low energy scenarios from the 1992 Supplement. Sources: Intergovernmental Panel on Climate Change (IPCC), 1990, 1992; Ausubel et al., 1988.
In a peculiar choice of words, the Intergovernmental Panel on Climate Change in 1990 designated as “Business-as-Usual” its scenario which stifled and even reversed the 130-year trend. “Business as Usual” was a scenario of technical regression. It essentially ignored the scientific and technical achievement of the past 300 years, including the achievements that make identification and estimation of the greenhouse effect possible. [End of excerpt.]
It’s good to see the lunatic BAU and 8.5 scenarios put in their proper place, if only after 35 years and a lot of harm. All this was in the open literature. Science is not always scientific.
Accurate, informative coverage of new Stoeckle-Ausubel paper on eDNA of New York’s East River
National Geographic How many rats are in New York City? Scientists find new clues in the East River. By Melissa Hobson
BBC Wildlife Magazine East River DNA reveals secrets of life (and death) in New York | Discover Wildlife by Helen Pilcher
C&EN Environmental DNA brings the East River’s fish population into focus By Sarah Braner
Green Prophet Poop in the East River shows the city’s rat problem and what people like to eat By Karin Kloosterman
MSN Environmental DNA study reveals new insights into aquatic life by Co-Pilot from Phys.Org
BioEngineer, United States Environmental DNA from NYC’s East River Uncovers Insights into Local Human and Wildlife Populations
Le Figaro, France Fish, humans… and rats: what DNA monitoring in urban rivers reveals by Delphine Chayet
Deutsche Presse Agentur via WEB.de, Germany What rivers reveal: genetic material paints a picture of entire ecosystems
el Mundo, Spain New York’s East River ‘knows’ how many rats the city has or what its inhabitants are eating by Florencia Da Silva
Mirage News, Australia East River transforms into living biosensor PLOS news release
The release generated at least 35 online stories across at least 14 countries and in 6 languages, with an estimated potential reach of around 130 million, impressive for East River fishes!
Jesse video interview on enduring environmental realities
Texas geologist Scott Tinker interviews Jesse Ausubel on “Enduring Environmental Realities” for 20 minutes beginning at 6:25 of this recording of the annual Energy Future Forum of the National Center for Energy Analytics (NCEA). Iddo Wernick is a Fellow of the NCEA. The Forum was presented by RealClearEnergy in partnership with the U.S. Chamber of Commerce on Friday 1 May in Washington DC. The agenda and entire program are here.
East River DNA reveals fish, urban wildlife, and what New Yorkers eat

Mark and Jesse’s year-long study of East River eDNA “Biomonitoring in the Anthropocene: Urban estuary environmental DNA tracks marine fish, terrestrial wildlife, and human diet” is published today in PloS ONE. The findings suggest that urban waterways anywhere could become continuous biosensors, tracking biodiversity, habitat restoration outcomes, and human impacts in real time. Among the most novel results of the weekly East River sampling were genetic indicators of human food consumption, of rat and other terrestrial wildlife populations, and the discovery of newly abundant fish species, thought to reflect the success of habitat restoration efforts. Our lively press release is posted here and graphic highlights here.
Vintage talk on Power and Climate
We post Jesse’s talk on “Power and Climate” delivered to executives of the US electric power industry in 2019. The 30-page pdf includes text (13 pages) followed by 17 pages of figures. Some highlights were excerpted here in February 2025, which makes us think the talk is standing the tests of time and earns posting.
Leonardo DNA in Science magazine
Science magazine’s Richard Stone publishes a superb survey of the Leonardo da Vinci Project, The Real da Vinci Code, occasioned by the posting to BioRxiv of our big paper, Biological signatures of history: Examination of composite biomes and Y chromosome analysis from da Vinci-associated cultural artifacts.
Media coverage is occurring, a couple of items are linked below.
The Telegraph Leonardo da Vinci DNA discovery could uncover lost paintings By Sarah Knapton
The Times Leonardo da Vinci’s DNA ‘found on artwork’ By Rhys Blakely
Irish radio program Drivetime interview with Jesse (starts with 2 minutes of patter)
Dubai radio The Agenda with Georgia Talley (starts at 56 minutes)
PHE high school intern presents to international eDNA meeting in Japan
Arya Kambhampati, Manhattan high school senior mentored by PHE’s Mark Stoeckle, presented on 13 December remotely to the 8th Annual Meeting of The eDNA Society in Yamaguchi, Japan on using eDNA to measure vertebrate biodiversity on oyster reefs such as those now being encouraged in New York Harbor. Congratulations, Arya, on your first talk to a major scientific conference.


Leonardo Da Vinci DNA Project public website
The purpose of the Leonardo da Vinci DNA Project is to create insights into the life and work of Leonardo da Vinci through application of rapidly advancing tools in biology and anthropology in close association with expertise from history and the arts. The new detective technologies span genetics and genealogy, microbiology, physical anthropology, chemistry, hydraulics, and visualization. A principal means for insight would be a genomic profile of Leonardo. The Project was conceived in 2014 by Brunetto Chiarelli, Cesare Marchetti, and Jesse Ausubel with crucial help from Henry de Lumley…and finally opens a public website.
Mark & Jesse give NOAA Webinar on Urban estuary eDNA
On 19 November Mark Stoeckle and Jesse Ausubel presented the NOAA ‘Omics Webinar on “Urban estuary eDNA reports seasonal abundance of marine fish, and, thanks to wastewater, tracks local wildlife, land birds, household pets, and human diet.” A video of the Webinar, one hour including Q&A, is here in YouTube. Thanks to Katie Poser and Nicole Miller for hosting and organizing.
Abstract: Here we analyzed vertebrate eDNA in New York City’s East River, a rocky estuary channel difficult to survey with mechanical gear and subject to wastewater discharge. There was a 10-fold increase in local marine fish eDNA in summer and seasonal differences among taxa consistent with known phenology. Levels of other vertebrate eDNA—domesticated animal, non-fish wildlife, and dietary fish—were correlated with human eDNA levels, consistent with a shared wastewater source. Wastewater eDNA identified the commonest urban mammals, land birds, and household pets. Proportions of dietary animal eDNA in wastewater closely approximated proportions in national consumption statistics, opening a window into human diet assessment. Wastewater eDNA analysis added value in an urban estuary survey of marine fish eDNA.