Harvesting wild animals for sale as food is a large, mostly illegal business that threatens wild animal populations and puts humans at risk for exotic infections, witness the SARS outbreak in 2003. Regulations and treaties exist, but before these can be enforced, one needs to establish the species origin of bushmeat and other derived marketplace products. Here DNA can help. In 1 September 2009 Conservation Genetics (open access article) researchers from University of Colorado, Barnard College, and American Museum of Natural History describe DNA barcodes for 23 species of South American and Central African primates, ungulates, and reptiles regularly harvested for bushmeat. Equally important as the DNA sequences, Eaton and colleagues report high success (179/204 samples (87.7%)) with primer cocktails first developed for fish DNA barcoding by Ivanova et al 2007, demonstrating these can serve as universal vertebrate primer cocktails. Intraspecific variation was low (mean 0.24%) and differences among congeneric species was generally high (average 9.77%), making assignment to known species straightforward using either tree-based maximum likelihood or character methods.

This report is focused on documenting barcodes of bushmeat species, using well-identified vouchered specimens (1 vouchered specimen labeled as Melanosuchus niger (Black caiman) was found to be Caiman yacare (Yacare caiman). The researchers did test a handful of unknown or partially identified specimens; all with recoverable COI sequences could be assigned to known species in the data set using the tree-based or character methods as described. Remarkably, Eaton and colleagues were able to recover COI DNA from 1 of 5 leather goods, which had been impounded by the US Fish and Wildlife Service as likely of CITES species origin. This proved to be Crocodylus niloticus (Nile crocodile). Recovering DNA from leather suggests many unsuspected household items have legible DNA barcodes.
I only wish the research report could have included pictures–there is so much more we might learn. There is an AMNH webpage describing the project which has several interesting images, although these are unlabeled and not referred to by the text. Perhaps we need a “mash-up” utility into which one could insert a scientific paper, which then would pull in relevant material–images, maps, links. Along these lines, there is a very neat Encyclopedia of Life NameLink utility which automatically detects scientific names and inserts hyperlinks to relevant EOL pages–try it!
 The horse-chestnut leaf miner moth Cameraria ohridella (link to
The horse-chestnut leaf miner moth Cameraria ohridella (link to 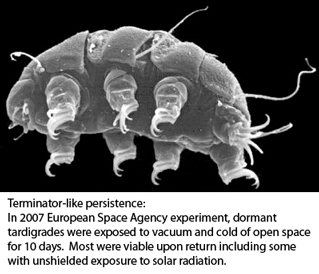 Tardigrades, commonly called water bears, are tiny (0.1-1.5 mm) water-dwelling invertebrates found in diverse environments. About 1000 species are known. Morphologic identification is difficult and may be limited to certain life stages–some species can be identified only from eggs, for example. Tardigrades can transform into a dormant state with remarkable ability to withstand extreme drying, cold, and radiation for prolonged periods, making them of interest for persons studying biology of tissue repair, aging and other fields.
Tardigrades, commonly called water bears, are tiny (0.1-1.5 mm) water-dwelling invertebrates found in diverse environments. About 1000 species are known. Morphologic identification is difficult and may be limited to certain life stages–some species can be identified only from eggs, for example. Tardigrades can transform into a dormant state with remarkable ability to withstand extreme drying, cold, and radiation for prolonged periods, making them of interest for persons studying biology of tissue repair, aging and other fields. “DNA barcoding involves sequencing a standard region of DNA as a tool for species identification. However, there has been no agreement on which region(s) should be used for barcoding land plants. To provide a community recommendation on a standard plant barcode, we have compared the performance of 7 leading candidate plastid DNA regions (atpF–atpH spacer, matK gene, rbcL gene, rpoB gene, rpoC1 gene, psbK–psbI spacer, and trnH–psbA spacer). Based on assessments of recoverability, sequence quality, and levels of species discrimination, we recommend the 2-locus combination of rbcL and matK as the plant barcode. This core 2-locus barcode will provide a universal framework for the routine use of DNA sequence data to identify specimens and contribute toward the discovery of overlooked species of land plants.”
“DNA barcoding involves sequencing a standard region of DNA as a tool for species identification. However, there has been no agreement on which region(s) should be used for barcoding land plants. To provide a community recommendation on a standard plant barcode, we have compared the performance of 7 leading candidate plastid DNA regions (atpF–atpH spacer, matK gene, rbcL gene, rpoB gene, rpoC1 gene, psbK–psbI spacer, and trnH–psbA spacer). Based on assessments of recoverability, sequence quality, and levels of species discrimination, we recommend the 2-locus combination of rbcL and matK as the plant barcode. This core 2-locus barcode will provide a universal framework for the routine use of DNA sequence data to identify specimens and contribute toward the discovery of overlooked species of land plants.”