The students of Niceville High School in the Florida Panhandle regularly contribute valuable samples from the sandy bottom of their beautiful shoreline to the near-shore (NaGISA) field project of the Census of Marine Life. On 23 January, Jesse had the privilege of visiting with the students and their exceptional instructor Rick Hernandez. The Okaloosa County School district reported the visit.
News
Distances and characters
Almost 4 years ago, in October 2005 Philos Trans R Soc Lond B Biol Sci researchers from American Museum of Natural History examined the then nascent DNA barcoding effort, looking at what methods were best for integrating the growing pool of DNA barcode data into systematics, the science of classifying organisms based on evolutionary history. Using real-world examples, authors DeSalle, Egan, and Siddall argued strongly for “characters” and against “distances” when using DNA barcode data to identify species, ie assigning specimens to known species and discovering new species. Of course, sequence data was already the backbone of modern systematics but it had primarily been applied to reconstructing evolutionary branching patterns (eg what pattern of divergences led to the various orders of birds) and less so to the definition of species. For example, most phylogenetic work included single examplars of each species. Analyzing sequence differences among and within closely-related species was more the domain of phylogeography which generally did not explicitly aim to define new species.
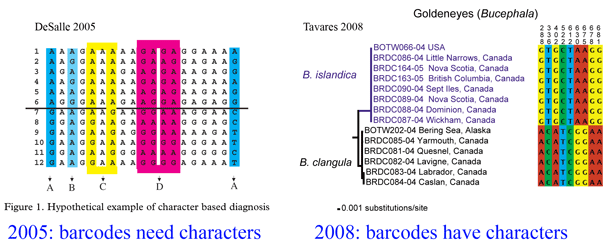 Here a brief aside. In analyzing sequences, “characters” refer to specific nucleotides (eg guanine (G) at position 138 in COI gene) and “distances” refer to per cent differences between sequences. So right away you can see that “characters” are intrinsic to the specimen’s DNA, whereas distances are defined only in relation to sequences from other specimens. Systematists like characters; for one, this enables integrating sequence and morphologic data. Characters are the grist for the computational workhorses of systematics, Parsimony and Maximum Likelihood. Meanwhile, beginning with the first paper published in 2003, distances displayed in neighbor-joining trees have been the usual heuristic approach for analyzing DNA barcode differences among and within species. A crucial advantage of neighbor-joining distance analysis is speed. Creating a NJ distance tree from 1000 648 bp barcode sequences might take a minute on a desktop computer whereas Maximum Likelihood reconstruction might take several weeks. Unlike reconstructing the Tree of Life, DNA barcoding is a recurrent exercise that repeatedly involves submitting new data from multiple known and unknown specimens, so a fast analytic method is essential.
Here a brief aside. In analyzing sequences, “characters” refer to specific nucleotides (eg guanine (G) at position 138 in COI gene) and “distances” refer to per cent differences between sequences. So right away you can see that “characters” are intrinsic to the specimen’s DNA, whereas distances are defined only in relation to sequences from other specimens. Systematists like characters; for one, this enables integrating sequence and morphologic data. Characters are the grist for the computational workhorses of systematics, Parsimony and Maximum Likelihood. Meanwhile, beginning with the first paper published in 2003, distances displayed in neighbor-joining trees have been the usual heuristic approach for analyzing DNA barcode differences among and within species. A crucial advantage of neighbor-joining distance analysis is speed. Creating a NJ distance tree from 1000 648 bp barcode sequences might take a minute on a desktop computer whereas Maximum Likelihood reconstruction might take several weeks. Unlike reconstructing the Tree of Life, DNA barcoding is a recurrent exercise that repeatedly involves submitting new data from multiple known and unknown specimens, so a fast analytic method is essential.
Four years later, where are we? Most DNA barcoding analyses continue to rely on NJ distance trees, and this approach has proven to be a durable heuristic, enabling one to distinguish among most species analyzed so far. Regarding species discovery, NJ distance trees demonstrate continued value as a first step in flagging divergent lineages that may represent new species. Here there is something of a roadblock, in that defining new species is a human judgement, sort of like a medical diagnosis, while sequences differences are like medical laboratory results. Community standards do not accept divergent mtDNA sequences as sufficient evidence to define a new species, although at the same time it is generally acknowledged that such sequences do indicate it is new, albeit one that hasn’t been officially defined yet. For example, in Nov 2008 news item researchers confidently assert “DNA tests identify new dolphin species,” (based on published article in Nov 2008 Mol Phylogenet Evol), yet include statement “it is awaiting a scientific name after a formal description.” I expect the researchers knew they had a new species with the first mtDNA sequence from a single individual! For DNA barcoding effort it should not be necessary to wait for final taxonomic decisions; we can proceed with publicly-disseminating a broad-range, fine-scale map of biodiversity, which can then be annotated with taxonomic information as it arrives. Like sky surveys and the human genome project, we should aim to make the “barcode biodiversity map” public as quickly as possible.
On the other side, it is now a commonplace observation that a 10X threshold (10 times the average intraspecific variation) is NOT a universal dividing line between intra- and inter-specific variation. To get technical, this was originally proposed as a screen for new species, but it has been taken as a dividing line between intra- and inter-specific distances, which it certainly is not; in the original 2004 paper (I am co-author) there are many sister species separated by distances less than the threshold. It has been a useful rhetorical target so maybe this issue won’t disappear just yet.
On the character front, there are more publications defining discriminatory DNA barcodes characters (eg Tavares and Baker 9 march 2008 BMC Evol Biol). It seems obvious to me that if, as is usually the case, sister species show large differences among and small differences within, then there must be diagnostic characters that distinguish them. The process of “translating” distances into characters should perhaps be a standard practice for nearest neighbor taxa in NJ trees; this would certainly give confidence (or not) as to whether one can reliably distinguish those species with less than 1% sequence difference. There is exciting development in character-based software tools (eg Ahrens et al 2007, Rosenberg 2007, Abdo and Golding 2007, Munch et al 2008) aimed at distinguishing the leaves (ie species) in addition to those already available for reconstructing the branches on the Tree of Life. I look forward to one that is friendly for non-specialists and works speedily on desktops!
Soy el arbol!!
In Chile in early January we visited some of the world’s largest tree plantations as well as the wondrous intact indigenous forest of Isla Mocha. Thanks to Savithri Narayanan for her photo of this sign “Soy el arbol!!” on a tree in a Chilean park with its true and poetic message (translated into English by Jesse).
“A reliable, consistent, and democratic tool for species discrimination”
 Human filariasis, caused by various species of insect-transmitted parasitic nematodes, affects more than 120 million persons in Africa, South America, and Southeast Asia, and includes elephantiasis and river blindness. In 7 january 2009 Frontiers Zool, 10 researchers from 5 institutions in Italy, France, Japan, and Venezuela apply DNA barcoding and traditional morphologic taxonomy to identification of parasitic filarioid worms. According to the authors, a molecular tool for identification of filiaria is a “desirable goal for many reasons” including “parasites conferred to diagnostic laboratories are often of poor quality due to the difficult[y] of sampling adults and undamaged organisms,” as a “method for the identification of filarioid nematodes in vectors,” and “nematode biodiversity is still highly underestimated both at the morphological and molecular level.”
Human filariasis, caused by various species of insect-transmitted parasitic nematodes, affects more than 120 million persons in Africa, South America, and Southeast Asia, and includes elephantiasis and river blindness. In 7 january 2009 Frontiers Zool, 10 researchers from 5 institutions in Italy, France, Japan, and Venezuela apply DNA barcoding and traditional morphologic taxonomy to identification of parasitic filarioid worms. According to the authors, a molecular tool for identification of filiaria is a “desirable goal for many reasons” including “parasites conferred to diagnostic laboratories are often of poor quality due to the difficult[y] of sampling adults and undamaged organisms,” as a “method for the identification of filarioid nematodes in vectors,” and “nematode biodiversity is still highly underestimated both at the morphological and molecular level.”
Ferri and colleagues analyze diagnostic utility of 12S and barcode-region COI sequences and morphologic examination by experts to an assemblage of data from 165 individual specimens (73 newly analyzed for this study) representing about 60 species. Their data set encompasses most of the important human and animal filarioid parasites, including Wuchereria bancrofti and Brugia malayi, agents of human tropical elephantiasis, Loa loa (human ocular filariasis), Onchocerca volvulus (human river blindness), and Dirofilaria immitis (dog and cat heartworm), plus specimens recovered from wild animals ranging from bats to toads.
The authors applied a medical test approach to the sequence data, looking at which distance cutoffs produced “minimum cumulative error,” in which they include type I false positive (failure to assign to correct species, analogous to oversplitting) and type II false negative (failure to distinguish between species; analogous to lumping). I find their approach refreshing in that it recognizes the uncertainty inherent in any identification method. Even “gold standard” tests have error rates. Just as a medical laboratory considers a range of factors when adopting a new test method–cost, speed, sensitivity, accuracy, replicability, and training requirements, for example, we might usefully look at methods for species identification, including traditional morphologic techniques, in a similar way. In taking such an approach, we can recognize there are often marked differences between the methods we use to detect something and the methods used to define it.
As a medical testing example, automated systems for rapid detection of bacteria in blood cultures rely on monitoring pressure changes in headspace gas in liquid culture bottles, as growing bacteria consume or produce gases. At the same time we do not define bacteria as “organisms that produce pressure changes in laboratory culture bottles,” for example. Similarly, percent differences between nucleotide sequences of the test specimen and those in a reference library might be a rapid way to “detect” a species, but this does not mean these are a defining characteristic of a species. We recognize species conceptually as independent evolutionary lineages, and practically on the basis of discriminatory characters (eg morphologic, behavioral, or nucleotide substitutions at specific sites). In the day-to-day work of specimen identification and detection of new species however, sequence distances may work just fine as diagnostic signatures.
Back to the article. Ferri and colleagues report COI worked better than 12S as a diagnostic, primarily due to difficulty in finding a consistent algorithm for aligning 12S sequences. With COI, the minimum cumulative error was 0.62% at a K2P distance threshold of 4.8%. The errors were due to low interspecific distances between 2 congeneric pairs [Onchocerca volvulus (human host) and O. ochengi (cattle); Cercopithifilaria longa (Japanese serow, a goat-antelope) and C. bulboidea (Sika deer); might some of the morphologic differences between these species pairs represent phenotypic changes induced by the different hosts?]. More sampling within species will help determine if it is possible to molecularly discriminate among these species using a character- rather than distance-based method.
The authors call for an integrated taxonomic approach to solve discrepancies between morphologic and molecular methods, and conclude “we propose DNA barcoding as a reliable, consistent, and democratic tool for species discrimination in routine identification of parasitic nematodes.”
DNA speeds discovery of overlooked species
 Just as new telescopes reveal previously hidden details of the universe, genetic surveys regularly reveal previously hidden (aka cryptic) species. Of course these species are cryptic only in the sense that morphological analysis is not the right tool to “see” them with. To my ear the word “cryptic” suggests camouflaged organisms that blend in with the environment, such as the Dead leaf butterfly Kallima inachus. Unlike camouflage, which is presumably a protection adaption, it is my impression there is nothing biologically special about morphologic crypsis except for the difficulty we have in recognizing it; that is, what we call cryptic species exhibit the same sorts of distinct ecological and behavioral adaptations found in those whose differences are more visible to the human eye.
Just as new telescopes reveal previously hidden details of the universe, genetic surveys regularly reveal previously hidden (aka cryptic) species. Of course these species are cryptic only in the sense that morphological analysis is not the right tool to “see” them with. To my ear the word “cryptic” suggests camouflaged organisms that blend in with the environment, such as the Dead leaf butterfly Kallima inachus. Unlike camouflage, which is presumably a protection adaption, it is my impression there is nothing biologically special about morphologic crypsis except for the difficulty we have in recognizing it; that is, what we call cryptic species exhibit the same sorts of distinct ecological and behavioral adaptations found in those whose differences are more visible to the human eye.
To restate the above, when multiple individuals are examined for gene(s) that reflect species-level differences (this is the essence of DNA barcoding), many animal and at least some plant species are discovered to be comprised of two or more genetic clusters, each carrying diagnostic nucleotide substitutions. When appropriate analytic tools are applied, these within-species clusters are often found to be reciprocally monophyletic lineages that have been reproductively isolated for hundreds of thousands to millions of years. In studies where the painstaking work of natural history observation has been carried out, these genetic clusters usually show ecological and behavioral differences and sometimes previously overlooked morphological distinctions, consistent with species-level status. In short, DNA analysis speeds discovery of new species. In many cases, it reveals species that would otherwise probably remain unrecognized indefinitely.
The premise of DNA barcoding is that a very short segment (ie for animals 648 bp COI barcode region) is usually sufficient to screen for new species and to assign specimens to known species. Of course, more sequencing is always of interest, but the added discriminatory value for detecting species-level differences is small compared to the added cost. Moving backwards in evolutionary time, a neighbor joining tree constructed with 648 bp barcode sequences often groups genera and families correctly; however it generally does not contain enough information to establish branching order or uncover deeper-level associations that are the heart of phylogenetic study, so there is plenty for systematists to do.
Now for some data. In 25 december 2008 Mol Phylo Evol researchers from University of Gothenburg and University of Florida report on Lumbriculus variegatus Muller, 1774, a segmented freshwater worm widely distributed in Europe and North America, commonly used as a model laboratory organism, in environmental toxicology, and sold as pet food for fish and amphibians under name “blackworm.” Part of the laboratory interest in L. variegatus lies in its remarkable ability to re-generate after fragmentation; any of the approximately 200 segments can re-form a complete adult worm; most populations reproduce through auto-fragmentation. Given that L. variegatus is a common, widely-distributed organism described over 200 years ago and is regularly used in scientific study, one might not expect any taxonomic surprises.
 Gustafsson and colleagues were initially studying a neuropeptide gene FMRFamide using L. variegatus purchased from a commercial supplier in California, with puzzling results suggesting polyploidy with multiple gene copies. This lead them to further characterize approximately 50 individuals collected at multiple sites in Europe and North America. Sequencing of COI, 16S, and ITS sorted the specimens into 2 phylogenetically distinct (maximum parsimony and Bayesian analysis) clades with 17% mean difference in COI, with the same genetic structure in mitochondrial COI/16S as nuclear ITS. Both clades were found in North America and Europe, sometimes at the same site. The authors conclude “it thus seems reasonable to regard these two main lineages within the L. variegatus complex as different species, regardless of which species concept one adheres to.” Of course, it may be they have rediscovered a named species; they caution that more study needs to be done including sampling the other named species in genus Lumbriculus (see EOL page).
Gustafsson and colleagues were initially studying a neuropeptide gene FMRFamide using L. variegatus purchased from a commercial supplier in California, with puzzling results suggesting polyploidy with multiple gene copies. This lead them to further characterize approximately 50 individuals collected at multiple sites in Europe and North America. Sequencing of COI, 16S, and ITS sorted the specimens into 2 phylogenetically distinct (maximum parsimony and Bayesian analysis) clades with 17% mean difference in COI, with the same genetic structure in mitochondrial COI/16S as nuclear ITS. Both clades were found in North America and Europe, sometimes at the same site. The authors conclude “it thus seems reasonable to regard these two main lineages within the L. variegatus complex as different species, regardless of which species concept one adheres to.” Of course, it may be they have rediscovered a named species; they caution that more study needs to be done including sampling the other named species in genus Lumbriculus (see EOL page).
DNA barcoding is an efficient instrument for revealing species-level differences. Routine application of DNA barcoding can enhance quality control in work with model organisms, cell lines, and collected specimens, and the long-term value of species descriptions.
WSJ Tucker
An opinion piece in the 29 December 2008 Wall Street Journal by William Tucker quotes Jesse’s low view of investments in so-called renewable energy sources from this interview in Weltwoche magazine.
Best wishes for 2009

The past year with the Barcode Blog has been exciting and challenging. Looking forward to 2009!
Mark Stoeckle
Program for the Human Environment
The Rockefeller University
Plant specialists work towards standardization
 In 26 November 2008 Mol Ecol researchers from University of British Columbia report on a meeting of 1200 plant specialists, entitled “Botany without Borders”, held on the campus in July 2008, which brought together the annual meetings of Botanical Society of America, the Canadian Botanical Association/L’Association Botanique du Canada, American Fern Society, and American Society of Plant Taxonomists. According to authors Kane and Cronk, DNA barcoding was a recurring theme of presentations and posters.
In 26 November 2008 Mol Ecol researchers from University of British Columbia report on a meeting of 1200 plant specialists, entitled “Botany without Borders”, held on the campus in July 2008, which brought together the annual meetings of Botanical Society of America, the Canadian Botanical Association/L’Association Botanique du Canada, American Fern Society, and American Society of Plant Taxonomists. According to authors Kane and Cronk, DNA barcoding was a recurring theme of presentations and posters.
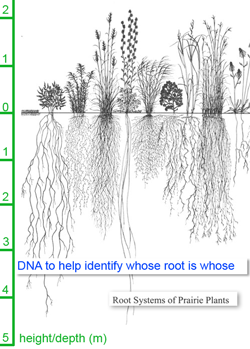
Plants continue to challenge a standardized approach to species identification using short DNA sequences from a uniform location on the genome, aka DNA barcoding. Genetic divergences among lineages make it difficult to design broad-range primers that amplify a desired target region across the diversity of plants and, at the same time, sequence differences among closely-related plant species are generally an order of magnitude fewer than those among animals, with the result that short sequences are often inadequate to assign specimens to species. Looking beyond these difficulties, the potential societal and scientific value of a standardized genetic identification method for plants is enormous. For one example cited in the meeting report, wild nutmeg trees of the genus Compsoneura can be identified by examining the tiny flowers on male trees, but trees are usually not in flower and female trees always lack these distinguishing characters. (It is remarkable that something as large as a tree can sometimes not be identified even by specialists!) In one study (Newmaster, Mol Ecol Notes 2007), a DNA barcoding approach using 2 short plastid sequences enabled identification of 94.7% of samples to species, compared to 40% using field characters. A standardized DNA-based approach should be a big boost to soil science by enabling the underground parts of plants, ie roots, to be readily named (Ridgway, BMC Ecol 2003).
 The authors conclude “DNA barcoding in plants is clearly here to stay and there is consequently an urgent need to rise to the scientific challenges it presents.” Some of those scientific challenges are explored in November 2008 Taxon by researchers from National Museum of Natural History, Washington, D.C., and National Center for Biotechnology Information, Bethesda, Maryland. Erickson and colleagues lay out a set of standard approaches to quantifying DNA barcoding success in plants.
The authors conclude “DNA barcoding in plants is clearly here to stay and there is consequently an urgent need to rise to the scientific challenges it presents.” Some of those scientific challenges are explored in November 2008 Taxon by researchers from National Museum of Natural History, Washington, D.C., and National Center for Biotechnology Information, Bethesda, Maryland. Erickson and colleagues lay out a set of standard approaches to quantifying DNA barcoding success in plants.
The authors state “PCR amplification must be the primary criterion for selecting a DNA barcode,” i.e. the chosen region should have the best rate of successful amplification across the diversity of plants. They suggest 90% or greater rate of recovery as a guideline. Second, they suggest each or any additional markers should improve PCR success by reducing the number of non-recovered PCRs by 50% and improve identification by at least 10%, using a parameter they call “probability of correct identification (PCI),” which is defined pretty much as it sounds. Applying this statistic to existing plant studies indicates the best results are with 2 plastid barcodes in which case PCI approaches an average of 90%, which of course includes much lower rates among some groups. Nonetheless, in local flora successful identification to species level may often approach 100%, because closely-related congeneric species are not present. The effort to establish a standardized genetic library of DNA barcodes for world’s plants is moving ahead.
mtDNA recovery from old bones hints at DNA durability, ubiquity
 In another seeming step towards Jurassic Park, two groups of researchers recovered full-length mitochondrial DNA sequences from 22,000 to 44,000 year-old bones of extinct European and North American bears. Full-length mtDNA has been recovered from similarly ancient specimens, but in those cases frozen tissues preserved in permafrost were used. Both groups used specialized PCR protocols employing several hundred primer pairs designed to recover short fragments, rather than one of the newer sequencing technologies, demonstrating the continued power of DNA amplification.
In another seeming step towards Jurassic Park, two groups of researchers recovered full-length mitochondrial DNA sequences from 22,000 to 44,000 year-old bones of extinct European and North American bears. Full-length mtDNA has been recovered from similarly ancient specimens, but in those cases frozen tissues preserved in permafrost were used. Both groups used specialized PCR protocols employing several hundred primer pairs designed to recover short fragments, rather than one of the newer sequencing technologies, demonstrating the continued power of DNA amplification.
In 28 july 2008 BMC Evol Biol Proc a group of 18 researchers led by Johannes Krause, Max Planck Institute, Germany, recovered full-length mtDNA from a 44,000 year old Ursus spelaeus (European cave bear) bone found in an Austrian cave, and from a 22,000 year-old skull of Arcdotus simus (American giant short-faced bear) from Eldorado Creek, Canada. In 11 november 2008 Proc Natl Acad Sci USA, 14 researchers led by Jean-Marc Elalouf, Institute de Biologie et Technologies de Saclay, France, report full-length U. spelaeus mitochondrial genome from a 32,000 year-old bone from the legendary Chauvet-Pont d’Arc Cave, home to the oldest rock art pictures ever found.
If we found a bone from one of these extinct bears in our backyard, could it be identified by its COI barcode? Submitting the long-ago bears’ COI barcode region sequences (positions 48 to 705) to BOLD ID engine flags both species as not in database, with a NJ tree similar to that created by full-length genomes (ie the extinct U. spelaeus is sister to U. arctos (Brown bear) and U. maritimus (Polar bear), and extinct Arcdotus simus is  sister to Tremarctos ornatus (Spectacled bear). Of course it would be difficult to recover a full-length sequence–what about the 130 base pair “mini barcode” proposed for broad-scale biodiversity analysis? This is within the size range(ie < 180 bp) that Elalouf and colleagues report best for recovery of ancient DNA. Remarkably, A. simus mini-barcode submitted to BOLD ID engine gives NJ tree correctly showing T. ornatus as its sister species and U. spelaeus mini-barcode correctly picks out U. arctos and U. maritimus as most closely-related species.
sister to Tremarctos ornatus (Spectacled bear). Of course it would be difficult to recover a full-length sequence–what about the 130 base pair “mini barcode” proposed for broad-scale biodiversity analysis? This is within the size range(ie < 180 bp) that Elalouf and colleagues report best for recovery of ancient DNA. Remarkably, A. simus mini-barcode submitted to BOLD ID engine gives NJ tree correctly showing T. ornatus as its sister species and U. spelaeus mini-barcode correctly picks out U. arctos and U. maritimus as most closely-related species.
Recovering DNA from ancient bones leads to CSI-like thoughts of where else we might usefully recover DNA for species identification. DNA has been recovered from naturally shed feathers, flakes of seal skin at breathing holes in polar ice, hair and saliva left by predators of sheep, bird faeces, and, turning to world of commerce, ancient and modern processed leather goods (Long 2007). I look forward to analyses of the many processed foods with what is currently an unverifiable “list of ingredients.”
Paul McCartney writes song for CoML Film
Beatle Paul McCartney’s blog confirms he is writing a song for the Jacques Perrin’s Ocean film made in cooperation with the Census of Marine Life.