“Amphibians are globally in decline, yet there is still a tremendous amount of u nrecognized diversity” observed Vences et al in 2005 Phil Trans R Soc B 360:1859, the first report applying DNA barcoding to amphibian diversity. Vences and colleagues highlighted the pressing need for fast and reliable identification tools, including for eggs and larva, which are often unrecognizable morphologically.
nrecognized diversity” observed Vences et al in 2005 Phil Trans R Soc B 360:1859, the first report applying DNA barcoding to amphibian diversity. Vences and colleagues highlighted the pressing need for fast and reliable identification tools, including for eggs and larva, which are often unrecognizable morphologically.
Here I focus on one technical aspect of DNA barcoding amphibians, namely designing primers that amplify the target sequence from a broad range of species. Previous research had shown remarkable mitochondrial sequence diversity among closely-related amphibians, and even within what appear to be single species, some of which may represent cryptic species. In the 2005 Proc R Soc B paper, researchers used COI primers designed for invertebrates (Folmer et al 1994); suprisingly these “worked in a large proportion of specimens”. They concluded “We support attempts to build up a global and complete cox1 database of [animal] eukaryotes”.
In 2005 Frontiers Zool 2:5 the same group of researchers quantified their PCR amplification success on specimens from 38 individuals representing 20 amphibian species. Using a well-established primer set for vertebrate 16s (Palumbi et al 1991) 38 of 38 (100%) samples amplified; with 3 COI primer sets (1 for invertebrates, 2 for birds), 36 of 38 (95%) amplified, although there was only 50-70% success for the individual COI primer pairs. The authors did not attempt to design new primers for amphibians. They concluded “we strongly advocate use of 16s rRNA as standard DNA marker for vertebrates to complement COI”. This seems reasonable but the advantages of standardizing on a single gene call for an effort to design primers that amplify COI from amphibians before abandoning the field to 16s or some other marker.
In 2007 Mol Ecol Notes Smith and colleagues from University of Guelph analyzed 83 amphibian COI sequences in GenBank to design new primers. The 3′ ends of the forward and reverse primers bind at 1st or 2nd codon position G-C residues, which they found to be highly conserved among amphibian species, and each primer contains three 2-fold degenerate sites. Using this set, they amplified full-length PCR products from 267 of 377 specimens (71%) representing 39 amphibian species (including Triturus vulgaris illustrated at right), and recovered an additional 34 sequences (9%) using a “mini-barcode” primer set designed for butterflies. The authors comment “many of the specimens…which failed to amplify had been fixed in formalin or were collected more than 15 years ago”, so further work to test these primers on fresh material and a diversity of species is needed.
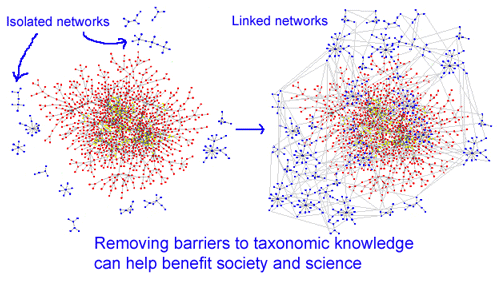
Amphibians are an exciting group. A comprehensive amphibian DNA barcode library will likely provide many, many new insights. I believe further work will help establish robust primer sets for amphibian COI sequences.
 Reference databases of DNA sequences used for species identification, ie DNA barcode libraries, are most powerful when the morphologic specimens are vouchered in a museum collection. This way, when there are puzzling results, DNA and morphologic specimens can be re-examined. However to date it has been challenging to recover DNA from small organisms without destroying them in the process.
Reference databases of DNA sequences used for species identification, ie DNA barcode libraries, are most powerful when the morphologic specimens are vouchered in a museum collection. This way, when there are puzzling results, DNA and morphologic specimens can be re-examined. However to date it has been challenging to recover DNA from small organisms without destroying them in the process. 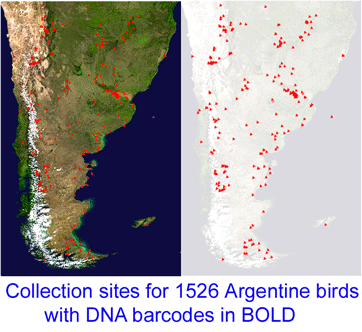 “This project, which started in December 2005, is a collaboration between MACN and the Biodiversity Insitute of Ontario/Canadian Center for DNA Barcoding (BIO/CCDB). In November 2006 the project was boosted by a grant from the Richard Lounsbery Foundation that supports expanded collecting efforts in Argentina, training of Argentine students at CCDB (2 trained so far), and establishment of a DNA laboratory at MACN.
“This project, which started in December 2005, is a collaboration between MACN and the Biodiversity Insitute of Ontario/Canadian Center for DNA Barcoding (BIO/CCDB). In November 2006 the project was boosted by a grant from the Richard Lounsbery Foundation that supports expanded collecting efforts in Argentina, training of Argentine students at CCDB (2 trained so far), and establishment of a DNA laboratory at MACN.  341 researchers from 44 countries gathered for the Second International Barcode of Life Conference, held at
341 researchers from 44 countries gathered for the Second International Barcode of Life Conference, held at