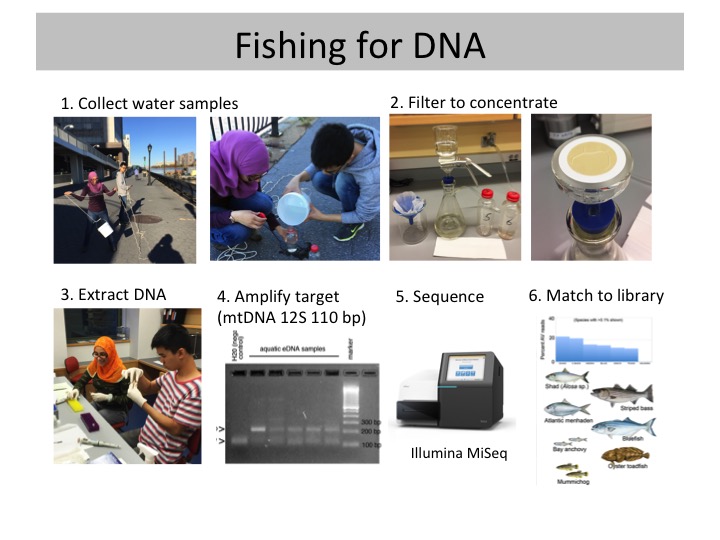
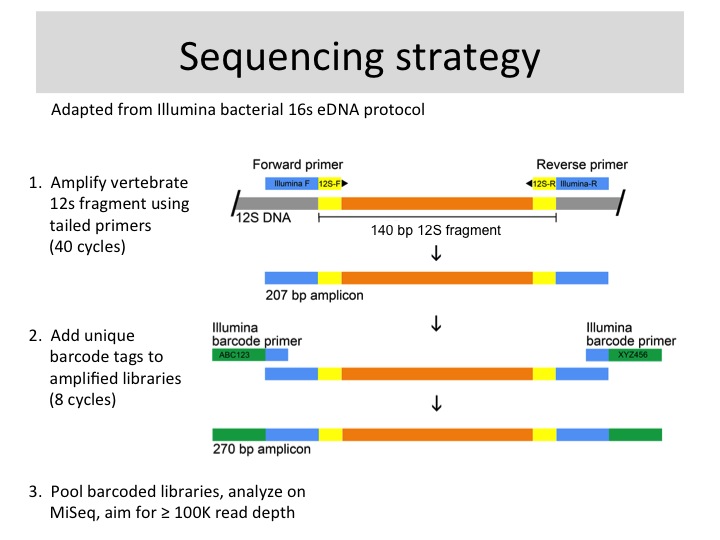
Methods
Link to NYC/NJ Fish eDNA home page
Download pdf NYC/NJ aquatic eDNA protocol (updated April 4, 2017)
Annotated references
Ficetola GF, Miaud C, Pompanon F, Taberlet P (2008) Species detection using environment DNA from water samples. Biol Lett 4:423-425 (open access link)
First demonstration of aquatic vertebrate eDNA detection; 15 mL water samples; ethanol ppt, spin, then extract DNA
Thomsen PF, Kielgast J, Iversen LL, Moller PR, Rasmussen M, Willerslev E (2012) Detection of diverse marine fish fauna using environmental DNA from seawater samples. PLOS ONE 7:e41732 (open access link)
Near shore marine fish along a pier; eDNA analysis equally or more effective than traditional methods; mostly used species-specific mtDNA primers; 30 x 50 mL samples pooled, then filtered 500 mL onto 47 mm diameter 0.45-µm pore size nylon filters (Osmonics, Penang, Malaysia); bead beating then Qiagen DNeasy
Thomasen PF, Kielgast J, Iversen LL, Wiuf C, Rasmussen M, Gilbert MTP, Orlando L, Willerslev E (2012) Monitoring endangered freshwater biodiversity using environmental DNA. Mol Ecol 21: 2565-2573 (link to abstract)
Detected endangered freshwater fish, frog, newt, river otter, and insects using species-specific qPCRs, also did 454 sequencing to confirm qPCR results; 3 x 15 mL samples per site, followed same ethanol ppt protocol as Ficetola 2008
Foote AD, Thomsen PF, Sveegaard S, Wahlberg M, Kielgast J, Kyhn LA, Salling AB, Galatius A, Orlando, Gilbert MTP (2012) Investigating the potential use of environmental DNA (eDNA) for genetic monitoring of marine mammals. PLOS ONE 7:e41781 (open access link)
Tested for dolphin DNA; 3/3 samples positive in or near captive dolphin pen; 1/8 samples positive near pen; 1/8 positive in open water sites where dolphins were nearby according to acoustic monitoring; also detected long-finned pilot whale; 3 x 15 mL each site; ethanol ppt, Qiagen extraction;
Kelly RP, Port JA, Yamahara KM, Crowder LB (2014) Using environmental DNA to census marine fish in a large mesocosm. PLOS ONE 9:e86175 (open access link)
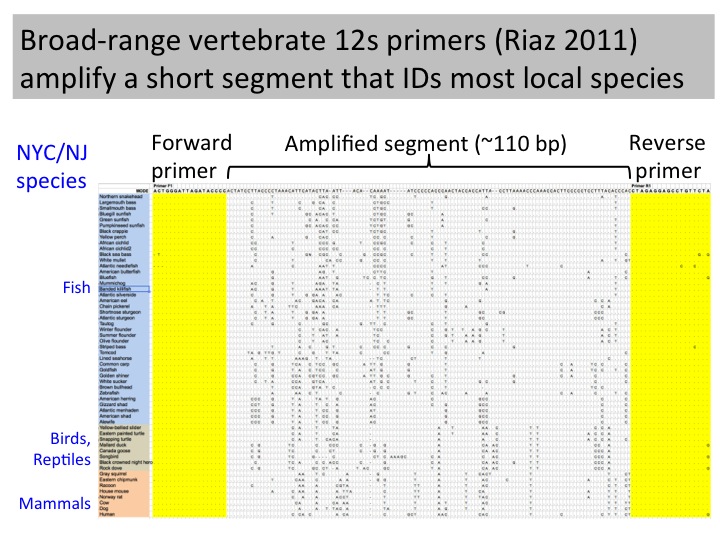
Aquatic eDNA detection applied to 4.5 million liter Monterey Bay Aquarium; 12s RNA primers amplifying 106 bp fragment detected all bony fish, needed other primers for cartilaginous fish and turtles; 1 L samples vacuum filtered onto 0.22-µm Durapore membrane filters (Millipore, MA, USA); Qiagen extraction
About the Bar Code of Life site
This web site is an outgrowth of
the Taxonomy, DNA, and Barcode of Life meeting held at Banbury
Center, Cold Spring Harbor Laboratory, September 9-12, 2003.
It is managed by Mark Stoeckle at the Program
for the Human Environment (PHE) at The Rockefeller University.
Contact: mark.stoeckle@rockefeller.edu
About the Program
for the Human Environment
The involvement of the Program for the Human Environment in DNA
barcoding dates to Jesse Ausubel's attendance in February 2002
at a conference in Nova Scotia organized by the Canadian Center
for Marine Biodiversity. At the conference, Paul Hebert
presented for the first time his concept of large-scale DNA
barcoding for species identification. Impressed by the
potential for this technology to address difficult challenges
in the Census of Marine Life, Jesse agreed with Paul on
encouraging a conference to explore the contribution
taxonomy and DNA could make to the Census as well as other large-scale
terrestrial efforts. In his capacity as a Program Director of
the Sloan Foundation, Jesse turned to the Banbury Conference
Center of Cold Spring Harbor Laboratory, whose leader Jan
Witkowski prepared a strong proposal to explore both the
scientific reliability of barcoding and the processes that
might bring it to broad application. Concurrently, PHE
researcher Mark Stoeckle began to work with the Hebert lab on
analytic studies of barcoding in birds. Our involvement in
barcoding now takes 3 forms: assisting the organizational
development of the Consortium for the Barcode of Life and the
Barcode of Life Initiative; contributing to the scientific
development of the field, especially by studies in birds, and
contributing to public understanding of the science and
technology of barcoding and its applications through improved
visualization techniques and preparation of brochures and other
broadly accessible means, including this website. While the
Sloan Foundation continues to support CBOL through a grant to
the Smithsonian Institution, it does not provide financial
support for barcoding research itself or support to the PHE for
its research in this field.