Evidence
What is the evidence that DNA barcoding is a reliable method for species identification?
For this commentary, “DNA barcoding” refers to nucleotide sequencing of PCR-amplified DNA corresponding to an approved barcode region, namely 5′ portion of COI for animals or rbcL + matK for land plants; and “species identification” refers to assigning the name of a known species to a specimen of unknown identity.
Acceptance by scientific community. For identification of known species, I think it is fair to say that DNA testing in general and DNA barcoding in particular are generally accepted in the scientific community as reliable methods. For example, the Canadian Centre for DNA Barcoding website has a compilation of peer-reviewed publications, which includes over 500 articles published since 2003. The primary limitation to identification is whether the relevant species and close relatives have yet been documented in the databases at the time they are queried. The BOLD database is strongest for multicellular animals (> 1,000,000 records as of May 2010; see chart), particularly arthropods and chordates. For plants, the general principles are the same, but so far there is much less documentation, as plant barcodes were not agreed-upon until last year (Hollingsworth et al PNAS May 2009), and there was not a large set of pre-existing data to 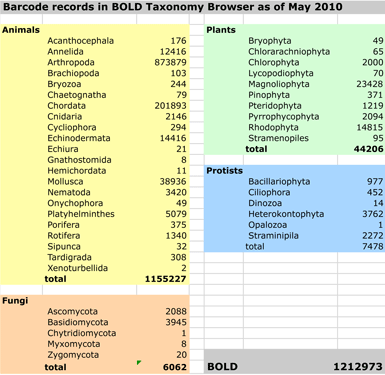 work with. Nonetheless, DNA barcoding of plants is ready for practical application and is providing immediately useful information (e.g. “DNA barcoding exposes a case of mistaken identity in the fern horticultural trade” Prior et al, Mol Ecol Resources April 2010) . For fungi, from perusing database it appears that ITS (internal transcribed spacer) and COI are informally accepted as barcodes. For protists and other domains of life, results so far suggest COI will serve as a primary barcode.
work with. Nonetheless, DNA barcoding of plants is ready for practical application and is providing immediately useful information (e.g. “DNA barcoding exposes a case of mistaken identity in the fern horticultural trade” Prior et al, Mol Ecol Resources April 2010) . For fungi, from perusing database it appears that ITS (internal transcribed spacer) and COI are informally accepted as barcodes. For protists and other domains of life, results so far suggest COI will serve as a primary barcode.
Most articles focus on DNA barcoding in a particular group and assess the accuracy of identification in that group. For example, in “DNA barcoding of commercially important salmon and trout species (Oncorhynchus and Salmo) from North America” (J Agricultural Food Chem 57:8379, 2009) Rasmussen and colleagues analyzed more than 1000 samples representing the 7 commercially important salmonid species from 143 sites across western North America including Alaska and Canada, (to capture possible variation within species) The authors found 100% separation of these species by DNA barcoding, i.e., distances among species were always greater than within species.
Forensic application. DNA barcoding for species identification has been used in legal cases (e.g. Cohen et al J Food Protection 72: 810, 2009). More general evidence is presented by Dawnay et al in “Validation of the barcoding gene COI for use in forensic genetic species identification” (Forensic Sci International 173:1, 2007). The authors conclude “this study demonstrates that the cytochrome c oxidase I gene enables accurate animal species identification where adequate reference sequence data exists.” As with any laboratory method, quality control and quality assurance (QA/QC) measures are essential (e.g. Morin et al J Heredity 101:1, 2010).
DNA barcode identification was designed to be a simple, straightforward method appropriate for wide use, and the results so far amply bear this out, including its use by high school students (e.g., “FDA pressured to combat rising ‘food fraud’,” Lyndsey Layton, Washington Post March 30, 2010). One aspect that needs work in my opinion are better explanations of the algorithms used for matching sequences to the databases and what the results mean. It still takes an expert to make sense of the data. Although the results are often obvious (e.g., 100% sequence identity to 10 barcode records of “Bos taurus (cow)”, interpretation is context dependent–a 100% match has a different meaning if a “neighboring” species differs by, say 1%, or if a congeneric species is not documented or is represented by a single record, for example. In my experience, identifications are usually straightforward, including recognizing ambiguous identifications. Nonetheless, for DNA barcoding to have the widest use, including in legal settings, it will be helpful to have better documentation of how we arrive at species diagnoses through DNA barcodes.
This entry was posted on Wednesday, May 26th, 2010 at 9:44 pm and is filed under General. You can follow any responses to this entry through the RSS 2.0 feed. Both comments and pings are currently closed.