 In forensic investigation, insect evidence helps date the time of death, as the various species that colonize corpses exhibit different stages of development according to time and temperature. Determining the post-mortem interval (PMI) rests on accurate species identification, including of immature forms. In Dec 2009 Int J Legal Med researchers from University of Wollongong, Australia, test DNA-based identification of Sarcophagidae flies, which lack distinguishing features as immature forms, and their adult identification requires “meticulous examination of subtle morphological differences, including regional hair presence and colour, body pigmentation and bristle length, placement and abundance”, and even then may need genitalic dissection for confirmation. As a result, sarcophagid flies are little used in forensic study, although being viviparous, they are “prospectively more reliable for PMI estimations compared with other initial dipteran colonisers” [the latter are mostly egg-laying species (e.g. callophorid blowflies), which hatch only if certain environmental conditions are met, adding uncertainty to PMI determinations].
In forensic investigation, insect evidence helps date the time of death, as the various species that colonize corpses exhibit different stages of development according to time and temperature. Determining the post-mortem interval (PMI) rests on accurate species identification, including of immature forms. In Dec 2009 Int J Legal Med researchers from University of Wollongong, Australia, test DNA-based identification of Sarcophagidae flies, which lack distinguishing features as immature forms, and their adult identification requires “meticulous examination of subtle morphological differences, including regional hair presence and colour, body pigmentation and bristle length, placement and abundance”, and even then may need genitalic dissection for confirmation. As a result, sarcophagid flies are little used in forensic study, although being viviparous, they are “prospectively more reliable for PMI estimations compared with other initial dipteran colonisers” [the latter are mostly egg-laying species (e.g. callophorid blowflies), which hatch only if certain environmental conditions are met, adding uncertainty to PMI determinations].
The researchers successfully recovered COI barcodes, without evidence of pseudogenes, from 85 adult specimens representing 16 species, using a single primer pair with degenerate bases previously applied to forensic blowflies (Nelson et al 2007 Med Vet Entomol). In NJ analysis, 14 of 16 species showed single clusters distinct from other species; the remaining 2 species showed deep divergences which the authors surmise may indicate cryptic species, perhaps more likely given that “taxonomic descriptions of the Australian Sarcophagidae have not been updated since the 1950s”.
Meikeljohn and colleagues demonstrate efficacy of COI barcodes as species-level identifiers for Australian sarcophagids. The tight intra-specific clustering in these flies appears identical to that seen in diverse animal groups including vertebrates, for example, yet flies are presumably several orders of magnitude more abundant. (As an aside, although the authors report their sequences and associated specimen data are deposited in BOLD, their data are not visible in “Public Projects”–I hope the authors will amend this.) What then limits mitochondrial variation within species? Or in the language of population genetics, why are effective population sizes for animal species uniformly small, unrelated to census population sizes? Like the nature of dark matter, explanation(s) await.
Addendum 11 Feb 2010: Dr. Meikeljohn reports that the sequences and associated data are scheduled to appear in BOLD and NCBI GenBank as soon article appears in print edition.
 In
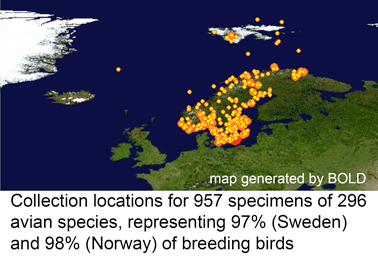
In 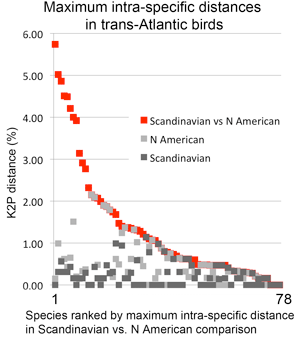 In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a
In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a  temporary placeholders is “taxon label”). As they note, there are many provisional names in GenBank (e.g. Ocyptamus sp. MZH S143_2004), so this is not a change in usual practice, except that the format of provisional names is standardized. As a starting point, Schindel and Miller propose a scheme developed by Council of the Heads of Australian Herbariums (CHAH) and recommend review by
temporary placeholders is “taxon label”). As they note, there are many provisional names in GenBank (e.g. Ocyptamus sp. MZH S143_2004), so this is not a change in usual practice, except that the format of provisional names is standardized. As a starting point, Schindel and Miller propose a scheme developed by Council of the Heads of Australian Herbariums (CHAH) and recommend review by 

 In
In  What lives in soil? In
What lives in soil? In 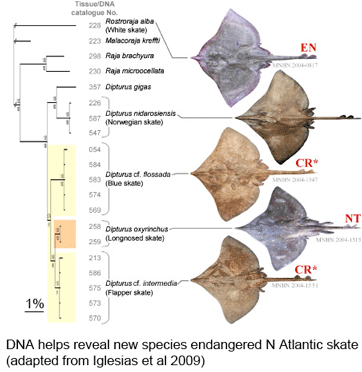 As the researchers report, this taxonomic oversight obscured the disappearance of one species, the flapper skate (D. cf. intermedia) because it was confused with the less threatened blue skate (D. cf. flossada). Iglesias and colleagues marketplace survey revealed additional sources of confusion. They analyzed 4,110 skates landed over a 2 year period from 103 fishing cruises in four main French ports by 41 different French commercial trawlers, and found that five skate species (included the two named above) from two genera are variously lumped together under just two marketplace names, the aforementioned “European common skate (D. batis)” and “longnose skate (D. oxyrinchus);” according to their analysis the latter species, formerly common, is also locally extirpated, and most specimens with this name represent other species.
As the researchers report, this taxonomic oversight obscured the disappearance of one species, the flapper skate (D. cf. intermedia) because it was confused with the less threatened blue skate (D. cf. flossada). Iglesias and colleagues marketplace survey revealed additional sources of confusion. They analyzed 4,110 skates landed over a 2 year period from 103 fishing cruises in four main French ports by 41 different French commercial trawlers, and found that five skate species (included the two named above) from two genera are variously lumped together under just two marketplace names, the aforementioned “European common skate (D. batis)” and “longnose skate (D. oxyrinchus);” according to their analysis the latter species, formerly common, is also locally extirpated, and most specimens with this name represent other species. Bluefin tuna are enormous (up to 15 ft/4.5 m, 680 kg/1500 lbs), high-speed (up to 54 km/h, as fast as racehorses) creatures that roam across oceans and return to ancestral waters to spawn. High demand has fueled intensive fishing by international fleets, resulting in 90% population declines heading towards extinction for all three species,
Bluefin tuna are enormous (up to 15 ft/4.5 m, 680 kg/1500 lbs), high-speed (up to 54 km/h, as fast as racehorses) creatures that roam across oceans and return to ancestral waters to spawn. High demand has fueled intensive fishing by international fleets, resulting in 90% population declines heading towards extinction for all three species, 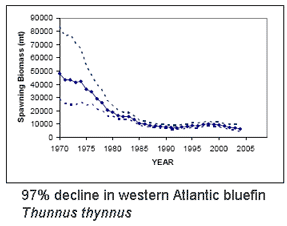 The eight species in genus Thunnus are not discriminated by regularly used nuclear loci and differ by about 1% or less in mitochondrial coding regions (e.g.,
The eight species in genus Thunnus are not discriminated by regularly used nuclear loci and differ by about 1% or less in mitochondrial coding regions (e.g.,  What hosts sustain arthropod disease vectors when they are not biting humans? In
What hosts sustain arthropod disease vectors when they are not biting humans? In