 Herbal products make a compelling case for DNA-based identification–how else to recognize dried bits of roots, leaves, stems, bark, and flowers from a multitude of species? In December 2009 J Nat Med, researchers from Ochanomizu University and Showa Pharmaceutical University, Japan, apply recently agreed-upon standards for DNA barcoding land plants, namely matK and rbcL, to distinguish among Dendrobium species. Dendrobium is a large (about 1200 species) genus of orchids widely distributed through east Asia to Philippines, Australia, and New Zealand. Over 50 Dendrobium species are used in traditional medicines and are thought to have various pharmacologic activities, although the active ingredient(s) are not yet characterized.
Herbal products make a compelling case for DNA-based identification–how else to recognize dried bits of roots, leaves, stems, bark, and flowers from a multitude of species? In December 2009 J Nat Med, researchers from Ochanomizu University and Showa Pharmaceutical University, Japan, apply recently agreed-upon standards for DNA barcoding land plants, namely matK and rbcL, to distinguish among Dendrobium species. Dendrobium is a large (about 1200 species) genus of orchids widely distributed through east Asia to Philippines, Australia, and New Zealand. Over 50 Dendrobium species are used in traditional medicines and are thought to have various pharmacologic activities, although the active ingredient(s) are not yet characterized.
Asahina and colleagues analyzed rbcL and matK from 12 samples representing 5 Dendrobium sp. and 3 hybrid cultivars whose genetic histories are uncertain. Single primer sets successfully amplified matK and rbcL from all specimens. The researchers cloned PCR products (and then sequenced at least 3 clones per species), rather than directly sequencing amplified products (rationale for the cloning step is not given). They found that matK, but not rbcL, distinguished among the five species; this is consistent with general observation that rbcL varies less among closely-related species than does matK. Results were similar when 22 matK Dendrobium sp. sequences from GenBank were added to analysis (bringing species total to 6), with one exception; 1 of 11 D. officinale GenBank matK sequences was unique, and in NJ diagram appeared on branch distant from the other 10. In this modest sampling, there was no intra-specific variation in the original 12 samples; some intra-specific differences were noted in 2 species in comparison with GenBank sequences.
 This study demonstrates advantages of DNA barcoding approach for plant identification. Of course, there is already a lot of interest in DNA identification of herbal plants in general and Dendrobium orchids in particular. For example, I found over a dozen articles describing DNA methods for distinguishing Dendrobium sp. However, the methods described are limited to identifying species in this one genus, which means one has to have a pretty good idea what the specimen is before applying DNA testing! This highlights the essential advantage of barcoding–a standardized approach can be applied to any unknown, and makes feasible creation of a comprehensive reference library.
This study demonstrates advantages of DNA barcoding approach for plant identification. Of course, there is already a lot of interest in DNA identification of herbal plants in general and Dendrobium orchids in particular. For example, I found over a dozen articles describing DNA methods for distinguishing Dendrobium sp. However, the methods described are limited to identifying species in this one genus, which means one has to have a pretty good idea what the specimen is before applying DNA testing! This highlights the essential advantage of barcoding–a standardized approach can be applied to any unknown, and makes feasible creation of a comprehensive reference library.
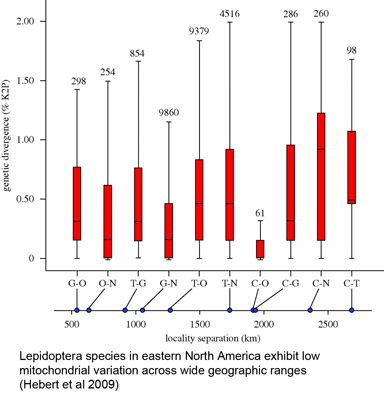
Looking ahead, we want to know more about intra- and inter-specific variation in plants. In animals, the patterning of mitochondrial variation is quite uniform, with intra-specific << inter-specific variation, such that most species form relatively tight clusters distinct from those of other species in NJ diagrams. Results so far in plants generally show little intraspecific variation in chloroplast genes (including rbcL and matK), but a diversity of distances among closely-related species. Assuming these early results are borne out, we then want to know why plants and animals differ? For more genetic variation in plants and animals, see Rieseberg et al Nature 2006, Fazekas et al Mol Ecol Res 2009).
 In forensic investigation, insect evidence helps date the time of death, as the various species that colonize corpses exhibit different stages of development according to time and temperature. Determining the post-mortem interval (PMI) rests on accurate species identification, including of immature forms. In
In forensic investigation, insect evidence helps date the time of death, as the various species that colonize corpses exhibit different stages of development according to time and temperature. Determining the post-mortem interval (PMI) rests on accurate species identification, including of immature forms. In 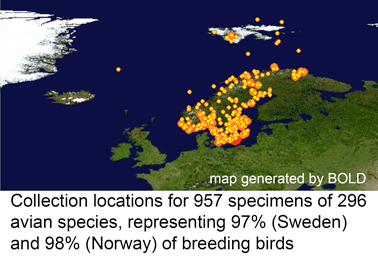 In
In 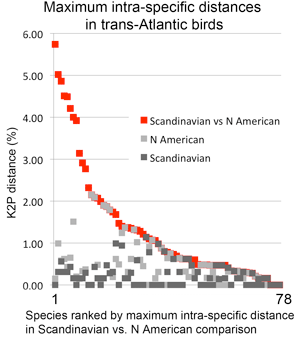 In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a
In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a  temporary placeholders is “taxon label”). As they note, there are many provisional names in GenBank (e.g. Ocyptamus sp. MZH S143_2004), so this is not a change in usual practice, except that the format of provisional names is standardized. As a starting point, Schindel and Miller propose a scheme developed by Council of the Heads of Australian Herbariums (CHAH) and recommend review by
temporary placeholders is “taxon label”). As they note, there are many provisional names in GenBank (e.g. Ocyptamus sp. MZH S143_2004), so this is not a change in usual practice, except that the format of provisional names is standardized. As a starting point, Schindel and Miller propose a scheme developed by Council of the Heads of Australian Herbariums (CHAH) and recommend review by 

 In
In