Results so far show most animal species correspond to clusters of closely-related mtDNA sequences, distinct from clusters of neighboring species. This patterning is so striking that if a neighbor-joining tree of mtDNA sequences were shown on the SAT, high school students could likely recognize the branches that correspond to species. For example, below is a nj tree of mtDNA sequences showing cryptic species of long-tailed shrew tenrecs in Madgascar (Olson et al 2004 Biol J Linnaean Soc 83:1). For an invertebrate example, see last week’s post.
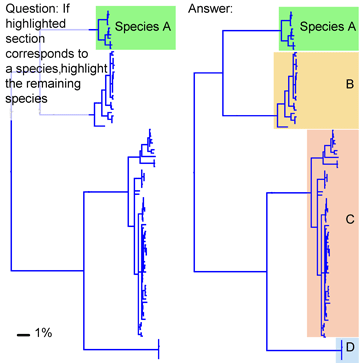
The remarkably widespread pattern of restricted intraspecific sequence variation in mtDNA in animals calls out for better scientific understanding. To my reading, most of the genetic taxonomic literature is focused on sequence differences because sequence differences are the necessary grist for reconstructing evolutionary history. Absence of variation means no characters, and no ability to generate evolutionary hypotheses. When absence of variation within species is found, it is often given the ad hoc and untestable explanation of being due to a recent population bottleneck. In individual cases this might seem plausible, but the hypothesis becomes absurd when applied to the large number of animal species that show low intraspecific variation. For example 97% of the 263 world cowrie species show constrained intraspecific variation (Meyer and Paulay 2005 PLoS Biol 3: e422); it is nonsensical to suppose all went through recent population bottlenecks. Low intraspecific variation is often said to indicate a small “effective population size”, but this is simply a restatement of the finding. As far as we know, many species are ancient and have enormous population sizes, factors that should permit variation. The burgeoning barcode libraries demonstrating limited intraspecific mtDNA variation in most animal species prompt the question, what erases history within species?
In April 2006 Science, researchers at Universite Montpellier, France (Bazin et al 2006 Science 312:570) report that population size does not influence mitochondrial diversity in animals, and hypothesize that mitochondrial DNA “probably undergoes frequent adaptive evolution”. Perhaps because this report threatens the foundation of many lines of research based on the assumption that mtDNA is a neutral marker whose diversity reflects population size, this study has elicited cautious commentary. Science’s own Perspectives piece concludes weakly “the diversity of mitochondrial DNA does not appear to reflect population size…and may be of only limited utility in understanding ecological, genetic, and evolutionary processes. It is ironic that the lack of recombination, once seen as a great asset of mitochondrial DNA, may be something of a problem in this context”. May be something of a problem? This study and the growing barcode surveys demonstrating limited mitochondrial sequence variation within most animal species overturn the assumptions of population biology and phylogeography and call for a new look at mitochondrial genetics.
This posting highlights one of the most interesting off-shoot findings to come out of the Barcode of Life Initiative. It’s no great surprise to find sequence divergence among species, even closely related and recently formed species. The surprise is how little mtDNA variation we’re finding within species. Early in the Barcode Initiative, we might have concluded that observed levels of intraspecific variation would increase as we obtained barcodes from more individuals and from different local populations, but in most cases that’s not the pattern that’s emerging.
The finding is certainly good news for barcoders, but as Mark’s blog points out, it raises an important question concerning the population dynamics of mtDNA. Eric Bazin gave a presentation at the Paris workshop organized by CBOL’s Data Analysis Working Group based on his recent Science article (see the workshop report at http://barcoding.si.edu/PDF/DAWG%20Paris%20Workshop%20Report%20-%2031%20July.pdf). The idea of “selective sweeps” is one conjecture. CBOL is definitely interested in promoting more work on this question!