Cool new barcode app
The US Global Positioning System (GPS), consisting of 24 to 32 satellites in medium earth orbit, cost $32 billion to develop and is supported by an annual budget of $1 billion. When the high resolution GPS signal was first made available to the public in May 2000 by President Bill Clinton, I imagine that few persons anticipated how useful it would be. Ten years later there are numerous, diverse applications, ranging from a smartphone app for finding the nearest post office in Australia to tracking animals across the Pacific. Like GPS, the Barcode of Life Database (BOLD) is a public, large-scale technology infrastructure resource. Similar to the trajectory with GPS, I expect that over the next 10 years BOLD will enable an expanding array of applications useful for students, consumers, commercial entities, regulators, researchers, and probably some just for fun.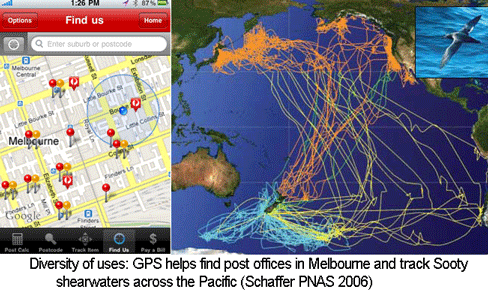
In November 2010 Molecular Ecology (request pdf from author) researchers from University of Guelph, Canada and Institut National de la Recherche Agronomique, France report on “molecular analysis of parasitoid linkages (MAPL)”. As background, parasitoid insects–many or most are wasps (order Hymenoptera)–lay eggs in the larvae of other insects, primarily Lepidoptera (butterflies and moths) and Diptera (flies). Host mortality may exceed 90%, and many parasitoids serve as useful biocontrol agents for agricultural pests. Parasitoid wasps are generally tiny and hard to distinguish morphologically, and identifying hosts may take years of patient observation. Recent molecular data show unexpected diversity and host specificity, i.e. many parasitoid species thought to be generalists are in fact comprised of multiple distinct lineages each limited to a single host.
In this study, Rougerie and colleagues looked at whether it was possible to identify the hosts by looking for leftover DNA in the abdomen of adult wasps. As an aside, the general approach in building up the barcode reference library for animals is to use broad-range primers that amplify COI from a wide taxonomic array of specimens. Now that parts of the library are established, it is possible to make use of the accumulated data to design primers that amplify specific taxonomic groups. Such taxon-restricted primers can help address interesting questions. In this study, researchers utilized two sets of primers, one set (primarily LepF1/LepR1) that amplified COI from the wasps and one set (LepF1/MLepR1) with a reverse primer that was specific to the potential hosts, namely Diptera and Lepidoptera. The first set successfully amplified COI from single legs of 297 adult wasp specimens thought to comprise more than 90 species and 20 genera. Using the same DNA extracts, the host-specific primers yielded PCR products from only 9 (3%) of these specimens, demonstrating good selectivity. Rougerie and colleagues then prepared DNA extracts from the abdominal segment of 3 species of hand-reared wasps (so that the host species were known), collected immediately after emergence. 29 (24%) of 120 specimens yielded readable PCR products, of which all except one matched to the known lepidopteran host species. The authors conclude that “MAPL has immediate applications in the agricultural sciences by facilitating selection of biological control agents” and that it “will drastically accelerate the registration of host-parasitoid associations and that the development of similar approaches for other orders of insects with complete metamorphosis will be equally productive.” I look forward to these new apps!
This entry was posted on Tuesday, December 14th, 2010 at 3:21 pm and is filed under General. You can follow any responses to this entry through the RSS 2.0 feed. Both comments and pings are currently closed.
December 14th, 2010 at 4:02 pm
This is a great series of posts with the Coulter notes and prior. THink about the progression of QR codes, a rarity in Europe and the Americas last year when we put them in Barcode art in Atlanta: http://www.ontariogenomics.ca/outreach/BOLD
Now ubiquitous in store windows, billboards, catalogs this holiday season. The app/handheld platform future is here and accelerating, including the Chrome OS and notebooks moving to assuming cloud functionality as the default. Barcorders to come. Keep up the great work!