Trans-Atlantic DNA survey reveals overlooked avian diversity in scientific heartland
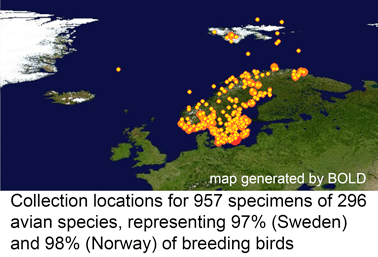 In January 2010 J Ornithol (open access article) researchers from Norway Natural History Museum, Swedish Museum of Natural History, University of Guelph, and Rockefeller University (myself) survey mitochondrial differences in 296 species representing 97-98% of Scandanavian breeding birds. 283 (95.6%) of species formed unique clusters; the remaining 13 species formed 5 clusters consisting of 2-4 species with shared or overlapping barcodes, which might reflect young species, hybridization with introgression, and/or a single gene pool. Surprisingly for such a relatively small geographic area, large sequence differences were found in 4 species, all of which have large breeding ranges that extend outside of Scandanavia; the authors propose these represent “a mixture of separate lineages that evolved in allopatry” and advise further sampling to “elucidate the phylogeographic history”.
In January 2010 J Ornithol (open access article) researchers from Norway Natural History Museum, Swedish Museum of Natural History, University of Guelph, and Rockefeller University (myself) survey mitochondrial differences in 296 species representing 97-98% of Scandanavian breeding birds. 283 (95.6%) of species formed unique clusters; the remaining 13 species formed 5 clusters consisting of 2-4 species with shared or overlapping barcodes, which might reflect young species, hybridization with introgression, and/or a single gene pool. Surprisingly for such a relatively small geographic area, large sequence differences were found in 4 species, all of which have large breeding ranges that extend outside of Scandanavia; the authors propose these represent “a mixture of separate lineages that evolved in allopatry” and advise further sampling to “elucidate the phylogeographic history”.
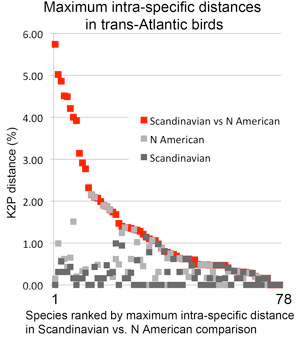
Johnsen et al take advantage of existing barcode library to compare species whose breeding ranges extend across the Atlantic. 19 (25%) of the 78 showed intercontinental divergences typical of species-level differences, including 8 species that had not been identified in prior work (data re-compiled in figure below). Most were inland species with discontinuous breeding ranges but there were unexpected exceptions such as Steller’s eider (Polysticta stelleri), which has what appears to be a continuous circumpolar breeding range. Three of the species formed paraphyletic clusters when combined with N American congeners, suggesting the inter-continental “conspecifics” are not even each other’s closest relatives.
 In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a world list of 8,616 species, which he judged to be “within 5 percent and certainly 10% of the final total”. The current IOC World Bird List v 2.3 recognizes 10,322 species (19% higher than Mayr’s estimate) and there is a steady stream of splits of existing forms, fueled by DNA sequence data. I believe DNA barcoding offers a way complete this process in a timely manner. If we analyzed multiple individuals from each of world’s named species, there would still be many areas of uncertainty, but at least the larger differences would be known. It is a scientific embarrassment that we are still discovering lineages that have been reproductively isolated for millions of years, in everyday birds no less!
In my view, this paper demonstrates that a survey approach produces a high level of discovery and hypothesis-generating, and leads me to question how well we understand diversity in birds, which are generally considered the taxonomically best-known large group of animals. Many of the species in the present study have been known to science for over 250 years, are resident in densely-settled, scientifically-advanced regions, and yet Johnsen and colleagues demonstrate hidden diversity. In 1946, Ernst Mayr compiled a world list of 8,616 species, which he judged to be “within 5 percent and certainly 10% of the final total”. The current IOC World Bird List v 2.3 recognizes 10,322 species (19% higher than Mayr’s estimate) and there is a steady stream of splits of existing forms, fueled by DNA sequence data. I believe DNA barcoding offers a way complete this process in a timely manner. If we analyzed multiple individuals from each of world’s named species, there would still be many areas of uncertainty, but at least the larger differences would be known. It is a scientific embarrassment that we are still discovering lineages that have been reproductively isolated for millions of years, in everyday birds no less!
There are over 300,000 avian tissue samples in the world’s museums, representing over 7,000 species (Stoeckle and Winker, Auk 2009). By my calculation, a modest number of these have been analyzed to date for species-level differences. For instance, by my count GenBank contains 13,361 cytochrome b sequences representing 4,320 avian species, and the All Birds Barcoding Initiative (ABBI) has so far collected 17,250 sequences representing 2,969 species. A concerted project of the world’s avian tissue collections employing DNA barcoding approach suggests an unmatched opportunity for large-scale, species-level genetics with many discoveries and hypothesis-generating findings which will inform various areas in evolutionary science. For instance, population genetics modeling starts with correctly identifying breeding populations (ie species). These samples may be eventually be analyzed in small batches, assuming they are not lost or destroyed, but the pace of standard research practices brings to mind the story of the Dead Sea Scrolls. Some were published soon after discovery in 1946, but the rest fell under the control of a committee of scholars and remained hidden not only from public but from other scholars for more than 40 years. When the monopoly was broken in 1991 (by researchers using a desktop computer to reconstruct texts from published concordances), some complained:
“Dr. Frank M. Cross, a scholar at the Harvard Divinity School who has worked with the scrolls since the 1950’s, said in a telephone interview that the publication of these unauthorized versions, which he described as “pirated,” would have no effect on the pace and publication schedules involving the actual scrolls. He defended his colleagues from the frequent charges of undue secrecy and procrastination, saying the critics did not understand the difficulties of working with the remaining unpublished documents that are mostly a collection of fragile fragments of parchment.” New York Times September 5, 1991
This entry was posted on Sunday, January 31st, 2010 at 10:20 pm and is filed under General. You can follow any responses to this entry through the RSS 2.0 feed. Both comments and pings are currently closed.