DNA helps reveal bat diets
June 9th, 2009
What do carnivorous animals eat? Predation drives evolution and underlies ecology, yet except for a few easily observed species, it is surprisingly hard to determine what eats what. In June 2009 Mol Ecol, researchers from University of Guelph and University of Western Ontario, Canada, apply DNA testing to help solve diet of Eastern red bat Laiurus borealis. L. borealis is the commonest tree-roosting bat in North America, ranging from Canada and United States east of the Rocky Mountains into Central and northern South America. Like other insectivorous bats, L. borealis uses echolocation to detect night-flying insects. Many moth species have evolved “ears” that detect the ultrasonic sounds emitted by bats and exhibit defensive behaviors in response to echolocation signals, making bats and moths an interesting study in predator-prey co-evolution.
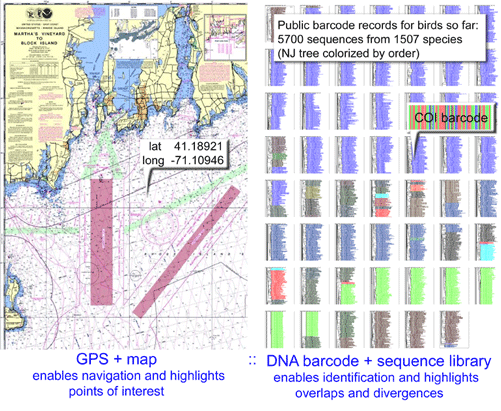
Clare and co-workers applied standardized DNA testing to insect parts in faecal samples collected from 56 mist-net trapped bats. Guano samples were frozen for up to 2 y then soaked in 95% ethanol for 12 h and examined with a dissecting microscope. Prey items including “legs, wings, antennae, eye cases, exoskeletal fragments, eggs” were isolated and stored separately in 96 well-plates. DNA extraction, amplification, and sequencing were performed using standard techniques and broad-range insect primers (LepF1/LepR1). COI sequences were compared to the 127,000 reference sequences of North American arthropods in BOLD database www.barcodinglife.org at the time of the study. Test sequences with >/=99% identity to reference sequence(s) and without equivalent similarity to other species in the database were given species-level identifications; those with less than 99% identity to reference sequence(s) were assigned to higher-level taxonomic categories.
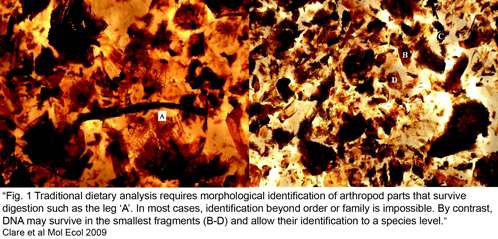 Clare et al obtained sequence data from 89% of 896 arthropod fragments; 78% of these were identified to species or genus level (the remaining 22% showed sequence similarity to bacteria, fungi, or were unidentifiable or chimeric), with a total of 127 prey species identified (125 insects, mainly lepidoptera including a number of economically important pest species, and 2 spiders). The “molecular scatology” approach documented greater diversity in prey species than prior studies based on morphologic analysis. Most prey were identified only once, with an average of 3.5 species per guano sample. Surprisingly, “more than 60% [of recovered insects] appear to have ears capable of hearing the echolocation hunting calls of L. borealis.” The authors speculate the abundance of eared moths might reflect bats hunting around streetlights, as moths in such brightly-lit environments are thought to use daytime predator-avoidance strategies rather than nocturnal responses to echolocation. There was a notable absence of actiid and tortricid moths, given their local abundance, suggesting these moths may have alternative predator-avoidance strategies.
Clare et al obtained sequence data from 89% of 896 arthropod fragments; 78% of these were identified to species or genus level (the remaining 22% showed sequence similarity to bacteria, fungi, or were unidentifiable or chimeric), with a total of 127 prey species identified (125 insects, mainly lepidoptera including a number of economically important pest species, and 2 spiders). The “molecular scatology” approach documented greater diversity in prey species than prior studies based on morphologic analysis. Most prey were identified only once, with an average of 3.5 species per guano sample. Surprisingly, “more than 60% [of recovered insects] appear to have ears capable of hearing the echolocation hunting calls of L. borealis.” The authors speculate the abundance of eared moths might reflect bats hunting around streetlights, as moths in such brightly-lit environments are thought to use daytime predator-avoidance strategies rather than nocturnal responses to echolocation. There was a notable absence of actiid and tortricid moths, given their local abundance, suggesting these moths may have alternative predator-avoidance strategies.
This study documents the diversity of L. borealis prey, and hints at how much more we will learn from broad application of standardized DNA analysis to food chains, including such unexpected findings as possible disruptive effects of man-made lighting on local ecosystems.
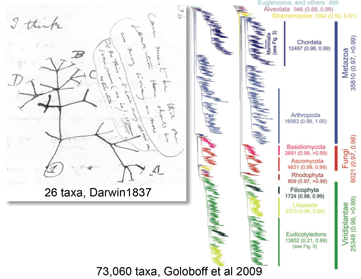

 Most DNA barcoding analyses look at DNA identification through the lens of established taxonomy, ie how well does sequence data capture the species-level taxonomic categories established by morphologic analysis? In the special issue article
Most DNA barcoding analyses look at DNA identification through the lens of established taxonomy, ie how well does sequence data capture the species-level taxonomic categories established by morphologic analysis? In the special issue article  Deep space telescopes gather light from the early universe, providing pictures of the unimaginably remote past. What about the biological universe–can we peer back in time? Geochemical evidence suggests life on Earth arose about 3.5 billion years ago and fossils reveal what life looked like as far back as 3.0 billion years, and important fossil discoveries across that whole span of time continue to be made. What about DNA? As Carl Woese first realized, DNA sequences of living organisms contain signatures of their evolutionary relationships, and enable reconstructing history as far back as the origin of replication, even before cells and DNA. At the near end of the time scale, recovery of DNA from historical samples can help identify organisms that lived hundreds, thousands, tens of thousands, or even, in a few cases so far, hundreds of thousands years ago.
Deep space telescopes gather light from the early universe, providing pictures of the unimaginably remote past. What about the biological universe–can we peer back in time? Geochemical evidence suggests life on Earth arose about 3.5 billion years ago and fossils reveal what life looked like as far back as 3.0 billion years, and important fossil discoveries across that whole span of time continue to be made. What about DNA? As Carl Woese first realized, DNA sequences of living organisms contain signatures of their evolutionary relationships, and enable reconstructing history as far back as the origin of replication, even before cells and DNA. At the near end of the time scale, recovery of DNA from historical samples can help identify organisms that lived hundreds, thousands, tens of thousands, or even, in a few cases so far, hundreds of thousands years ago. “Too often a journal’s decision to publish a paper is dominated by what the Editor/s think is interesting and will gain greater readership — both of which are subjective judgments and lead to decisions which are frustrating and delay the publication of your work. PLoS ONE will rigorously peer-review your submissions and publish all papers that are judged to be technically sound. Judgments about the importance of any particular paper are then made after publication by the readership (who are the most qualified to determine what is of interest to them).”
“Too often a journal’s decision to publish a paper is dominated by what the Editor/s think is interesting and will gain greater readership — both of which are subjective judgments and lead to decisions which are frustrating and delay the publication of your work. PLoS ONE will rigorously peer-review your submissions and publish all papers that are judged to be technically sound. Judgments about the importance of any particular paper are then made after publication by the readership (who are the most qualified to determine what is of interest to them).”

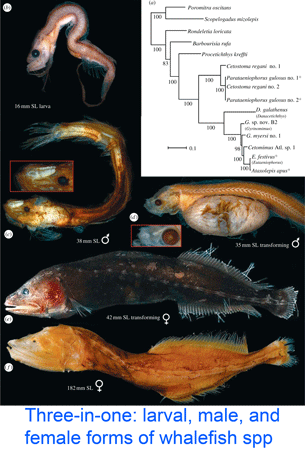 DNA helps flag genetically divergent forms that may represent cryptic species and is equally valuable the other way around: in linking morphologically diverse forms that occur within species. In
DNA helps flag genetically divergent forms that may represent cryptic species and is equally valuable the other way around: in linking morphologically diverse forms that occur within species. In  Mitochondria are the power plants of the cell, consuming oxygen and breakdown products of sugars, amino acids, and fatty acids to produce energy as ATP and heat. As originally proposed by
Mitochondria are the power plants of the cell, consuming oxygen and breakdown products of sugars, amino acids, and fatty acids to produce energy as ATP and heat. As originally proposed by 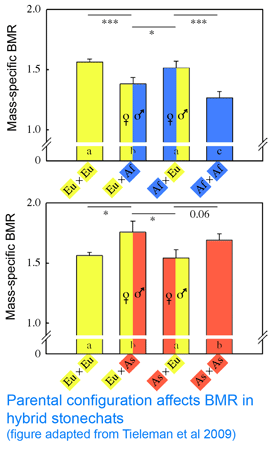 In
In