In current Aquatic Conserv Marine Freshwater Ecosys, researchers from Muséum national d’Histoire naturelle, France, report on 80 years of taxonomic confusion that has contributed to near extinction for a once abundant north Atlantic skate. Iglésias and colleagues found that two forms, lumped together in 1926 as European common skate (Dipturus batis, Linnaeus 1798), in fact represent distinct species with morphologic, genetic (in mitochondrial genome), and life history differences.
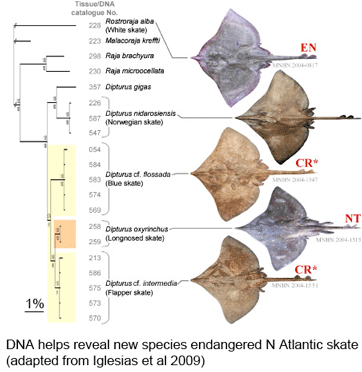 As the researchers report, this taxonomic oversight obscured the disappearance of one species, the flapper skate (D. cf. intermedia) because it was confused with the less threatened blue skate (D. cf. flossada). Iglesias and colleagues marketplace survey revealed additional sources of confusion. They analyzed 4,110 skates landed over a 2 year period from 103 fishing cruises in four main French ports by 41 different French commercial trawlers, and found that five skate species (included the two named above) from two genera are variously lumped together under just two marketplace names, the aforementioned “European common skate (D. batis)” and “longnose skate (D. oxyrinchus);” according to their analysis the latter species, formerly common, is also locally extirpated, and most specimens with this name represent other species.
As the researchers report, this taxonomic oversight obscured the disappearance of one species, the flapper skate (D. cf. intermedia) because it was confused with the less threatened blue skate (D. cf. flossada). Iglesias and colleagues marketplace survey revealed additional sources of confusion. They analyzed 4,110 skates landed over a 2 year period from 103 fishing cruises in four main French ports by 41 different French commercial trawlers, and found that five skate species (included the two named above) from two genera are variously lumped together under just two marketplace names, the aforementioned “European common skate (D. batis)” and “longnose skate (D. oxyrinchus);” according to their analysis the latter species, formerly common, is also locally extirpated, and most specimens with this name represent other species.
For newly rediscovered blue and flapper skates, the researchers report 20 diagnostic substitutions in approximately 2600-nucleotide segment spanning 12s and 16s RNA. Other than the 10 mitochondrial sequences included in this report, I find only two other D. batis sequences in GenBank (and none as yet under either of two resurrected names). It is remarkable that so little genetic information has been collected for such recently abundant, commercially important (annual landings in 1000’s of tons), and now threatened species. To aid standardized application of molecular identification techniques, I hope the authors will also analyze COI barcode sequences for their specimens. Then I look forward to school children aiding conservation and helping find new species by DNA barcoding specimens from their local fish markets!
It is indeed a pity that COI was not used in conjunction with this study as more than 150 Dipturus specimens representing more than 30 species have been barcoded and their sequence profiles and collateral data are lodged in the Barcode of Life Data system
(see: http://www.barcodinglife.com/views/taxbrowser.php?taxid=53054)
Moreover, COI barcode sequences have even been included in the description of a new species of Dipturus in order to help disambiguate and reconcile the application of names going forward…
(see: http://www.mapress.com/zootaxa/2008/f/z01921p046f.pdf)
yet we cannot place the results of the French study within the larger international sampling because they did not include COI. Note that if they would send samples of their taxa, FISH-BOL could cover the expense of barcoding so that their results can be nested within the larger global survey.